In translational biomarker research, improving specificity often increases assay complexity—particularly in immunoassay development1. We recently explored this tradeoff using a novel bioluminescent, multi-epitope assay designed to detect Alzheimer’s disease (AD)-relevant phosphorylated tau (pTau) species in blood. Despite the inherent challenges of plasma as a matrix, the assay showed strong concordance with amyloid PET imaging in a feasibility study, suggesting that certain conformational tau epitopes could serve as peripheral indicators of central amyloid pathology2.
At the core of the platform is a ternary split-NanoLuc® luciferase system—a proximity-driven luminescence approach specifically engineered for translational use in complex biological matrices. Unlike binary complementation systems that require precise alignment of luciferase fragments, the ternary design incorporates a scaffold protein that passively organizes the system (Figure 1). Luminescence is generated only when multiple tagged antibody fragments bind spatially adjacent epitopes on the same target, enabling signal generation that is driven by analyte recognition rather than nonspecific molecular crowding.
To mitigate high background in protein-rich matrices, the luciferase components were mutationally optimized to reduce spontaneous reassembly, improving baseline suppression and signal-to-noise performance. The system was empirically refined across a variety of sample types—including plasma, serum, saliva, nasopharyngeal swabs, urine, and potable water—and is compatible with point-of-care and high-throughput formats, supporting its translational robustness.
In our exploratory AD study, plasma samples were pre-analytically processed (VeraBindTM) and interrogated using a three-epitope pTau detection strategy. This assay requires all three epitopes to be present on the same pathologically relevant form of pTau, ehancing specificity for disease-associated isoforms and reducing detection of benign or fragmented tau species (Figure 1). The resulting luminescent signal was clean and sharply resolved, enabling the application of a potential binary threshold to stratify samples. This approach may help reduce the interpretive ambiguity seen in some existing pTau assays, where a significant proportion of cases fall into an indeterminate range. In this small feasibility cohort, the assay yielded an AUC of 0.93 and 88% concordance with amyloid PET imaging, highlighting its potential utility in translational research contexts3.
Overall, this study illustrates how proximity-based bioluminescent detection platforms, when combined with sample conditioning and epitope-driven capture strategies, may support the development of more accessible and interpretable assays for complex biomarkers in challenging matrices.
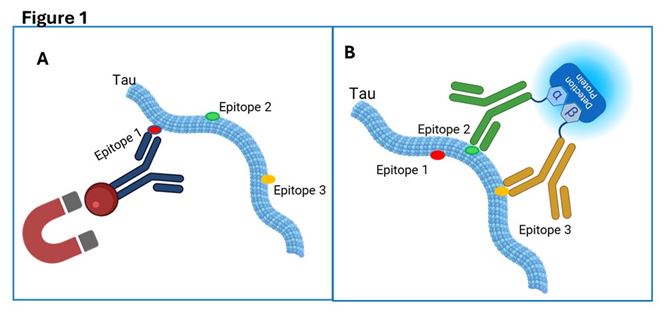
Figure 1: Schematic of a three-epitope-dependent immunoassay using ternary split-NanoLuc® luciferase system.
This novel multi-epitope immunoassay begins with capturing the target analyte via epitope 1 using a magnetic bead-conjugated antibody (A), allowing physical separation of bound versus unbound analyte. Detection only occurs if the analyte also presents epitopes 2 and 3, which are recognized by two distinct antibodies. These antibodies are each conjugated to complementary peptides (alpha and beta) of the ternary-split NanoLuc® luciferase enzyme that reconstitutes active enzyme only when both antibodies are co-localized on the same captured molecule (B). This three-epitope requirement enhances assay specificity and conformational fidelity.
References
- Findlay, J.W., et al., Validation of immunoassays for bioanalysis: a pharmaceutical industry perspective. J Pharm Biomed Anal, 2000. 21(6): p. 1249–73.
- Salvadó, G., et al., Plasma Phosphorylated Tau 217 to Identify Preclinical Alzheimer Disease. JAMA Neurol, 2025
- Soldo J, Ansari K, Torio E, Ressler V, Brouette C, and Dart M (2025). A Novel Bioluminescent, Multi-Epitope Assay to Detect Alzheimer's Disease-Specific Tau Consistent with Alzheimer's Amyloidosis. Association for Diagnostics & Laboratory Medicine Annual Meeting 2025, Chicago, Il; July 2025.


